My work sits at the intersection of statistical modeling, numerical methods, and software engineering. The common thread is building robust systems that turn noisy, high-dimensional data into quantitative conclusions through deep learning, Bayesian inference, or scalable data infrastructure, depending on the problem.
Co-founder and CTO of NoRD Bio Inc .
Publication Spotlight
(full
list indexed on
Google Scholar)
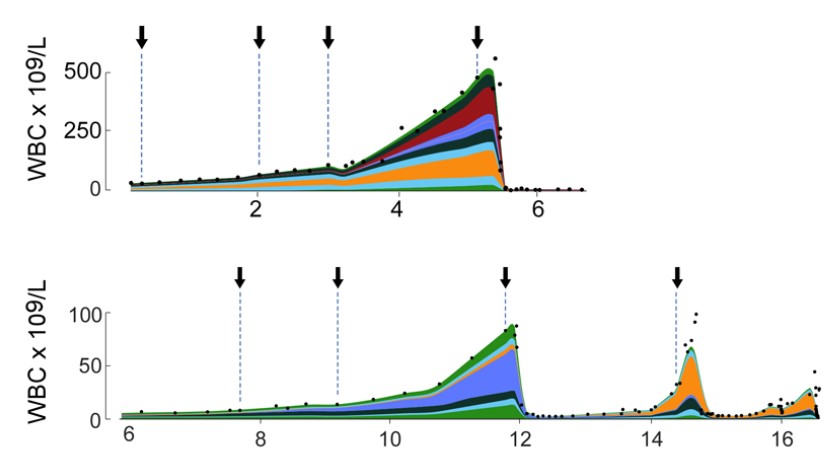 |
Growth dynamics in naturally progressing chronic
lymphocytic leukaemia *
We analysed the indolent growth
of CLL by reconstruction of the diseased cell population
from DNA sequencing data. This analysis was made
possible by our Bayesian clustering tool
PhylogicNDT.
|
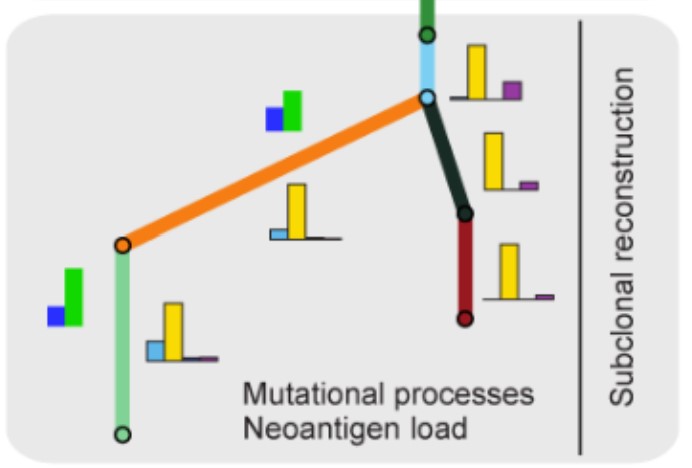 |
Comprehensive analysis of tumour initiation, spatial and temporal progression under multiple lines of treatment *
This paper is a comprehensive presentation of
PhylogicNDT.
We used it to reconstruct the trajectory of tumor growth in lung adenocarcinoma.
While every feature is individually showcased in other,
high
profile
peer-reviewed publications, this remains the best reference.
|
* co-first author
Selected Projects
Deep Learning for Molecular Diagnostics
At NoRD Bio I designed the computational framework behind a novel sequencing-based diagnostic platform. The core challenge was detecting extremely rare variant signals in high-dimensional sequencing data. I trained deep and probabilistic ML models to classify observations, and built a statistical framework for error modeling across millions of sequencing reads. Infrastructure is fully cloud-based (AWS), defined as code with Terraform, and run through CI/CD pipelines to maintain reproducibility between research iterations and production.
Bayesian Optimal Experimental Design
The statistical thinking behind that diagnostic work has deeper roots. My doctoral thesis at Columbia focused on Bayesian experimental design for nonlinear dynamical systems, where the goal was to select the most informative experiments before running them, reducing the total number needed to identify a model. I developed a framework combining Laplace approximation with automatic differentiation to evaluate expected information gain across candidate designs. For parameter inference in nonlinear hierarchical models with population heterogeneity, I implemented Sequential Monte Carlo samplers. The outer loop selects experiments that maximize information gain; the inner loop fits ODE-based kinetic models to time-series data, performs model criticism, and updates beliefs. I also built a Bayesian model selection layer to compare competing model structures (e.g., distinguishing reaction mechanisms) rather than just fitting parameters within a single model.
PhylogicNDT: Bayesian Clustering of Cancer Evolution
Before Columbia, I applied a similar Bayesian perspective to cancer genomics. At the Broad Institute I was a core developer of PhylogicNDT, a suite of tools for reconstructing the evolutionary history of tumors from DNA sequencing data. My main contributions were a fully custom Dirichlet process mixture model for identifying subclonal cell populations, and a tree-structured sampler for inferring the latent phylogenetic relationships between those populations. Both components are written in Python and were open-sourced. I also developed a filter to remove alignment artifacts from sequencing data, which was accepted by the consortium and applied to all datasets. The tools have been used in dozens of high profile, published studies.
Large-Scale Genomic Data Infrastructure
Building good models is only part of the problem; they need infrastructure that can run reliably at scale. Across roles at the Broad Institute and Ennov1, I built and optimized pipelines for processing large genomic datasets. At the Broad, this meant designing cloud workflows on GCP for whole-genome and whole-exome sequencing data, including parallelizing legacy serial code and building automated quality control checks. At Ennov1, I worked on a clinical genomics platform where each workflow run processed over 1TB of data through a Kubernetes cluster. I profiled resource utilization systematically, tuned container allocations, and restructured pipeline stages to optimize both cost and wall-clock run time. I also built a reporting microservice that generated and delivered patient reports to a customer-facing frontend.
Real-Time Object Detection on Edge Hardware
Not all inference happens in the cloud. I also ported trained object-detection models to a TensorRT and CUDA inference stack for deployment on edge devices. The work involved writing custom CUDA kernels, converting model graphs to TensorRT-optimized representations, and conducting systematic profiling with NVIDIA Nsight to identify and remove bottlenecks. The goal was to achieve real-time inference throughput for a computer vision application.
You can connect with me on LinkedIn.
I am a proud member of iCons alumni
network at UMass Amherst.
Thank you
iCons faculty and alumni for making this program possible!